Cloud Computation and Storage
Introduction
OmicsBox uses OmicsCloud, BioBam's Scientific Cloud Computing Platform, to execute many bioinformatics tools in a secure and scalable environment. OmicsCloud is an AWS-based system designed for high-performance computing (HPC), optimized for demanding bioinformatics algorithms. Users can run tools and pipelines without setup or maintenance, enabling scalable and parallelized cloud data analysis.
OmicsBox also provides Cloud Storage services that allow users to store data directly in the cloud, saving local disk space and enabling easy data access and sharing among colleagues.
Cloud Computation
Cloud Units
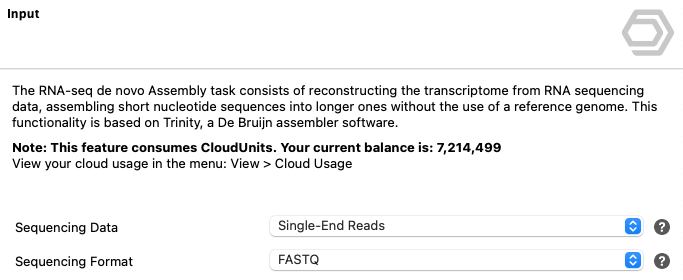
OmicsBox offers many tools that run in the cloud. These operations incur costs for CPU usage, data storage, and transfer, which are measured in Cloud Units. Cloud Unit consumption reflects computational resource usage such as CPU time and data volume. The demand for resources varies across tools depending on their algorithms, input data, and user-selected parameters.
Most tools can be used at no additional charge with an active subscription.
Currently, OmicsBox charges Cloud Units only for sequence Alignment and Assembly features (see list below). Consumption for these features is based on the CPU time (in seconds) required to complete the task. All other tools are included in the subscription at no extra cost.
- Functional Annotation Module
- Alignments: BLAST, Custom Database BLAST (also used via KEGG), Diamond, and InterProScan
- Genome Analysis Module
- Assembly: ABySS, SPAdes, Flye
- Alignments: BWA, Bowtie
- Transcriptomics Module
- Assembly: Trinity
- Alignments: STAR, BWA
- Metagenomics Module
- Assembly: MEGAHIT, metaSPAdes
Figure 1. Cloud Units balance overview in OmicsBox.
Cloud Sync
Cloud Sync is the asynchronous execution model used by many OmicsBox tools. When a tool runs with Cloud Sync, the computation takes place entirely in the cloud, independent of the local OmicsBox application. This provides several key advantages:
- Run without keeping OmicsBox open. After submitting a Cloud Sync job, OmicsBox can be closed or even the computer can be shut down. The analysis continues running in the cloud.
- Results saved in cloud storage. Output files are written directly to the user's Cloud Files space. When OmicsBox is reopened, the results are automatically available.
- Cloud files as input. Cloud Sync tools can read input data from Cloud Files, avoiding the need to upload files before each run. This is especially useful for large datasets or when chaining multiple analysis steps.
- Real-time monitoring. While a job is running, its status, progress, and log messages can be monitored from the Cloud Usage view.
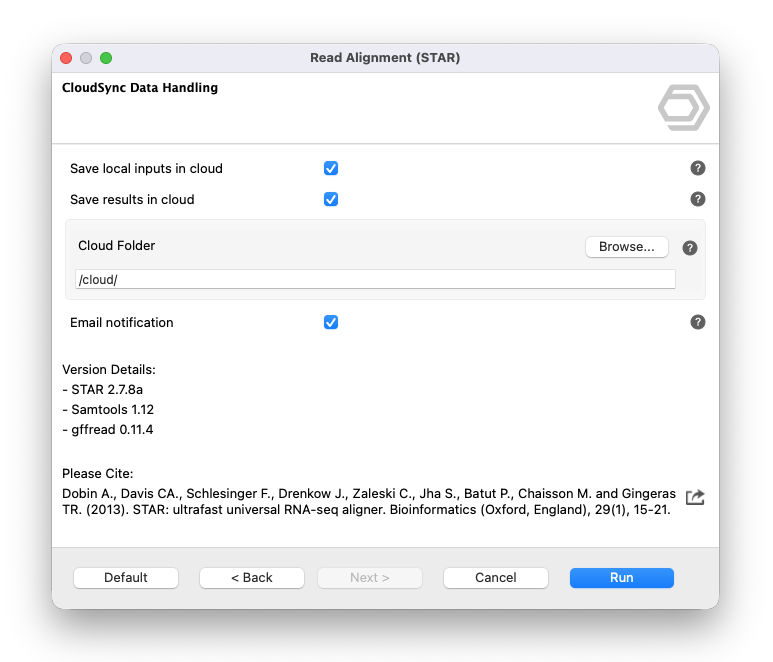
Cloud Sync is available for an increasing number of tools across all modules. Tools that support Cloud Sync display a CloudSync Data Handling page in their wizard (Figure 2), allowing users to configure where input files are stored and where output files are saved.
The CloudSync Data Handling wizard page provides the following options:
- Save local inputs in cloud: uploads local input files to the cloud before the analysis starts.
- Save results in cloud: stores output files in the user's Cloud Files space after the analysis completes.
- Cloud Folder: specifies the cloud directory where input and output files are stored.
- Email notification: sends an email when the job finishes.
Figure 2. CloudSync Data Handling page in the STAR Read Alignment wizard.
Note
Cloud Sync jobs use the same Cloud Units billing as other cloud tools. Most tools are included in the subscription. Only Alignment and Assembly tools consume additional Cloud Units based on CPU time.
Cloud Files
The Cloud Files view displays the files stored in the user's cloud space. Open it from the menu View > Cloud Files.
Every user has an individual cloud storage area. The context menu provides options to manage files. You can use drag and drop to copy files or folders from the local computer or the Local Files tab into the cloud.
Share files
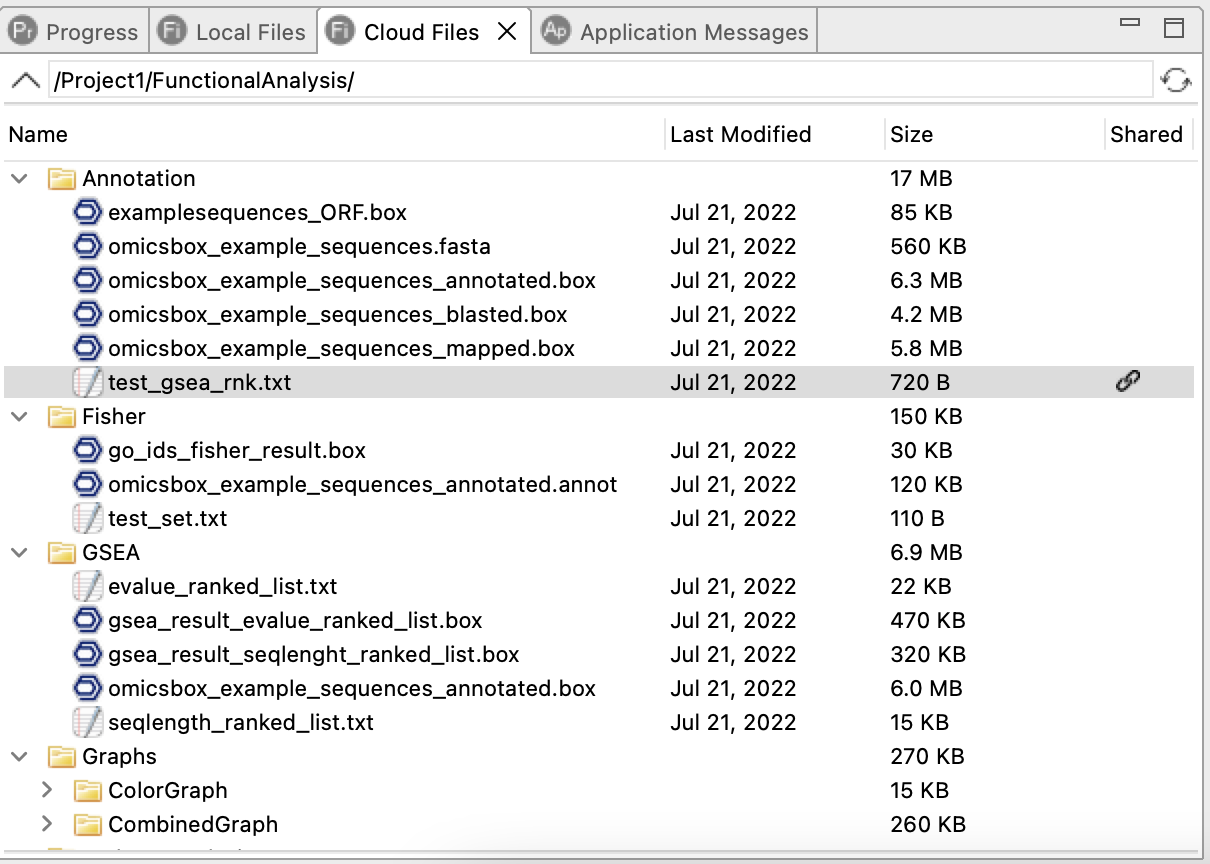
The Share option in the Cloud Files context menu allows sharing private files with others through a link. When a file is shared, a link icon appears in the Shared column. Anyone with the link can download the file using a web browser.
If a file is already shared, the Copy Shared Link menu option copies the existing link. The Unshare option makes a file private again. After unsharing and resharing, the new link will be different from the previous one.
Current limitations of the sharing functionality:
- Only individual files can be shared, not directories.
- Files are shared via link. Sharing with a specific user is not supported.
Figure 3. Cloud Files view showing the user's cloud storage.
Figure 4. Context menu with sharing options.
Figure 5. Shared link indicator in the Cloud Files view.
Review Cloud Usage
The Cloud Usage view provides a complete overview of cloud computation history, storage costs, and job monitoring. Open it from the menu View > Cloud Usage.
Summary statistics
The top section of the Cloud Usage view displays 3 key metrics:
- Used Storage: the amount of data stored in the user's cloud space.
- Estimated Monthly Cost: the projected Cloud Units to be charged by the end of the month based on current storage. The first 5 GB are free for each user every month, and data cost is measured daily.
- Available Cloud Units: the remaining balance of Cloud Units in the account.
For more details on pricing and Cloud Units consumption, visit the Cloud Computation page on the BioBam website.
Cloud Usage table
The main section of the view contains a table listing all cloud jobs and transactions. The table includes the following columns:
| Column | Description |
|---|---|
| Status | Current state of the job: Running, Done, Waiting, Sending, Cancelled, or Error. |
| Options | Action icons for interacting with the job (see below). |
| Submitted | Date and time when the job was submitted. |
| ID | A user-assigned identifier for the job. |
| Task | The name of the tool or service that was executed. |
| Progress | Completion percentage for running jobs. Shows 100% for finished jobs. |
| Runtime | Elapsed time from submission to completion, or current elapsed time for running jobs. |
| Input | Size of the input data sent to the cloud. |
| Output | Size of the output data produced by the job. |
| Consumption | Total Cloud Units consumed by the job. |
| Charged | Cloud Units actually billed. Tools included in the subscription show "0 (included)". |
The table supports sorting by any column, filtering by date ranges, and text search. Data can be exported to a tab-separated file using the view menu.
Options column
The Options column provides quick actions for each job through a set of icons:
- Stop: cancels a running job. Only visible while the job is still in progress.
- Log Viewer: opens a dedicated view with the cloud execution logs for the job. Logs can be exported to a text file using the view menu of the log panel.
- Messages: opens the progress messages view, showing the detailed algorithm messages produced during execution.
- Details: opens a task details viewer that displays the full job metadata, including parameters, timestamps, and resource usage.
- Cloud Files: navigates to the Cloud Files view and opens the output folder for the job, providing direct access to the result files stored in the cloud.
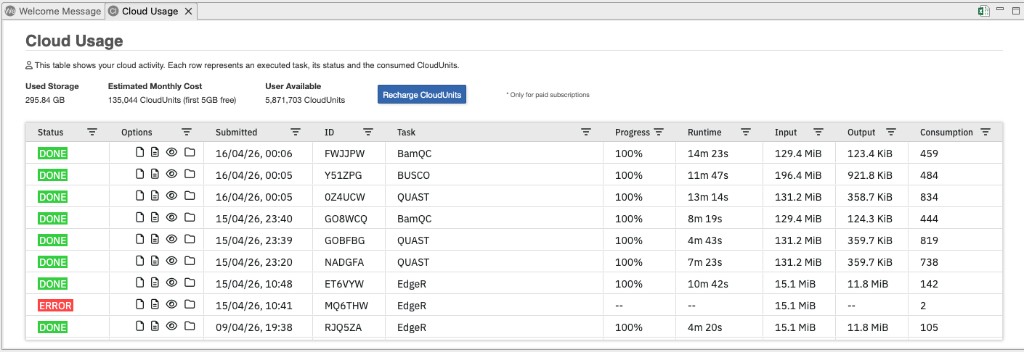
Figure 6. Cloud Usage view showing job history with status, options, progress, runtime, and resource consumption.